Of course they did, you reply.
But not so fast!
There has, of course, been a lot of interest lately in tracing one’s DNA to find out one’s individual ancestry. But I haven’t seen much about the results of doing that for an entire ethnic group, or set of ethnic groups.
So I just had to wade through a recent study in the journal Cell exploring the ancestry of populations in the Middle East. It wasn’t easy reading, at least for someone like me with minimal background in DNA, genome sequencing, and words like “polygenic.” (I did take a college course in genetics and animal behavior, but we’re talking the mid-’60s, so today’s world of genetics and DNA bears little resemblance to what I had learned, even if I could somehow remember what I had learned!)
Anyhow, recent events in Afghanistan have of course prompted interest in questions like “What language do they speak in Afghanistan?” Apparently most Americans have no idea and invent answers like “Afghan” or “Arabic.” But Afghan isn’t a language, and Afghanis aren’t Arabs, so those answers won’t work, even though the mistake is understandable. (The correct answer is that they primarily speak Dari and Pashto; Dari is just the dialect of Persian spoken there, basically the same as the Farsi spoken in Iran, and Pashto is an older form of Persian; there are no significant number of Arabs in the mix.)
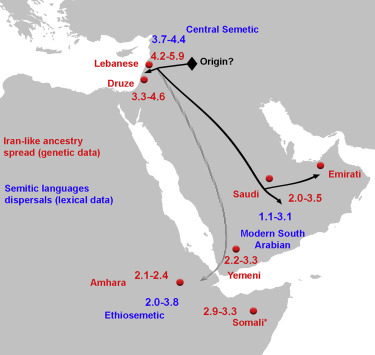
All of this just gives you a context for the map shown above, which gives a visual representation of the dispersal of Iran-like and Semitic languages in the region. Clearly linguistic and ethnic aspects are closely related (linguistics, in fact, used to be part of anthropology). But what does all this have to do with DNA and the human genome? There’s no way I can competently summarize the article in question, so I am just going to quote their abstract verbatim:
The Middle East region is important to understand human evolution and migrations but is underrepresented in genomic studies. Here, we generated 137 high-coverage physically phased genome sequences from eight Middle Eastern populations using linked-read sequencing. We found no genetic traces of early expansions out-of-Africa in present-day populations but found Arabians have elevated Basal Eurasian ancestry that dilutes their Neanderthal ancestry. Population sizes within the region started diverging 15–20 kya, when Levantines expanded while Arabians maintained smaller populations that derived ancestry from local hunter-gatherers. Arabians suffered a population bottleneck around the aridification of Arabia 6 kya, while Levantines had a distinct bottleneck overlapping the 4.2 kya aridification event. We found an association be- tween movement and admixture of populations in the region and the spread of Semitic languages. Finally, we identify variants that show evidence of selection, including polygenic selection. Our results provide detailed insights into the genomic and selective histories of the Middle East.
Categories: Life, Linguistics, Teaching & Learning